Find out more about our ambitious upgrade project, delivering more brightness, more coherence, and greater speed of analysis to UK science. More about Diamond-II
![]()
Find out more about Diamond's response to virus research.
![]()
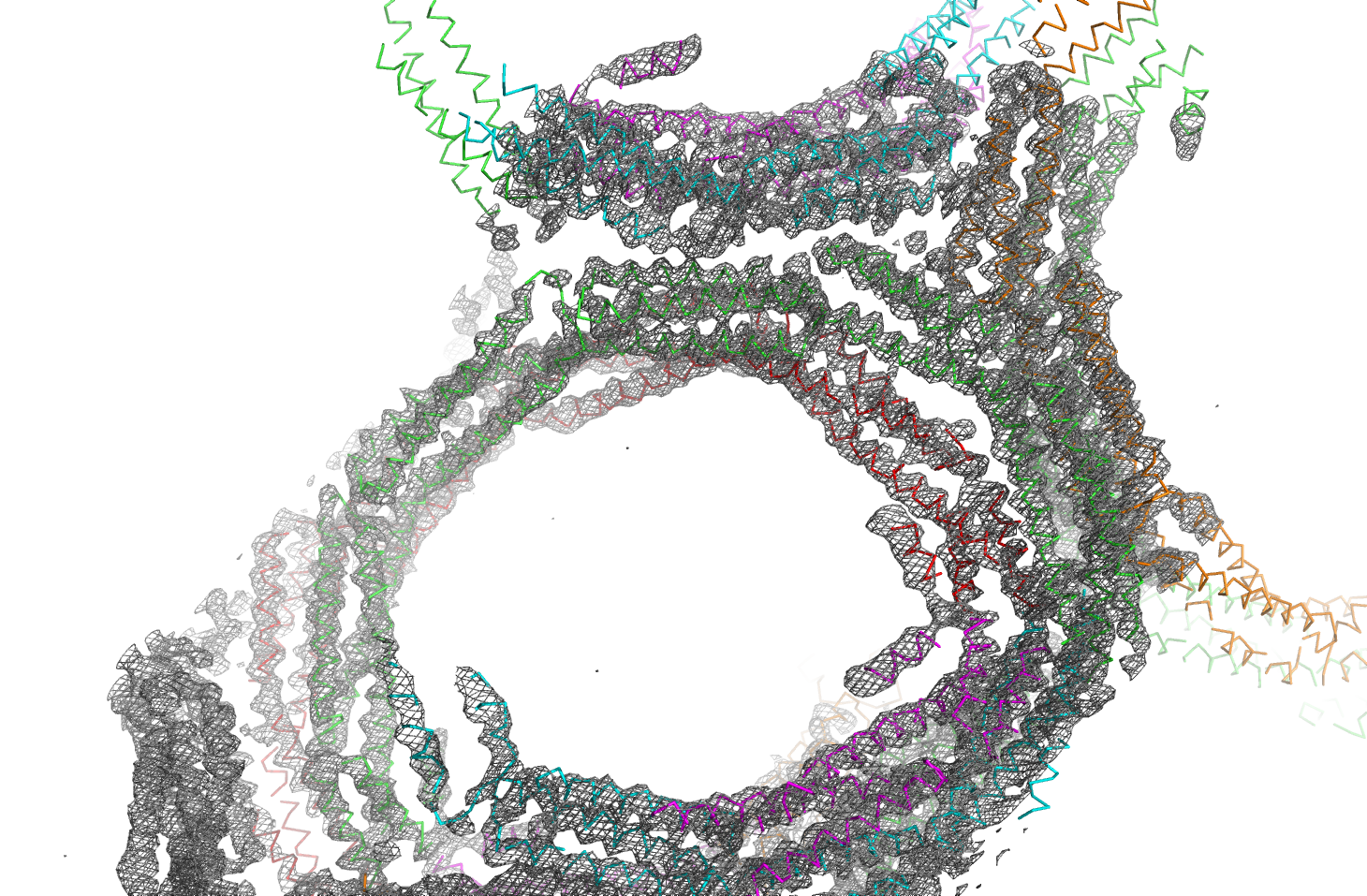

Scientists have uncovered important new information about how bacteria grow and multiply, potentially leading to the discovery of much-needed new antibiotic drugs. Using Diamond Light Source – the UK’s synchrotron science facility – scientists have determined the structure of EzrA: a protein that helps bacteria to build their defensive cell walls as they divide and spread.
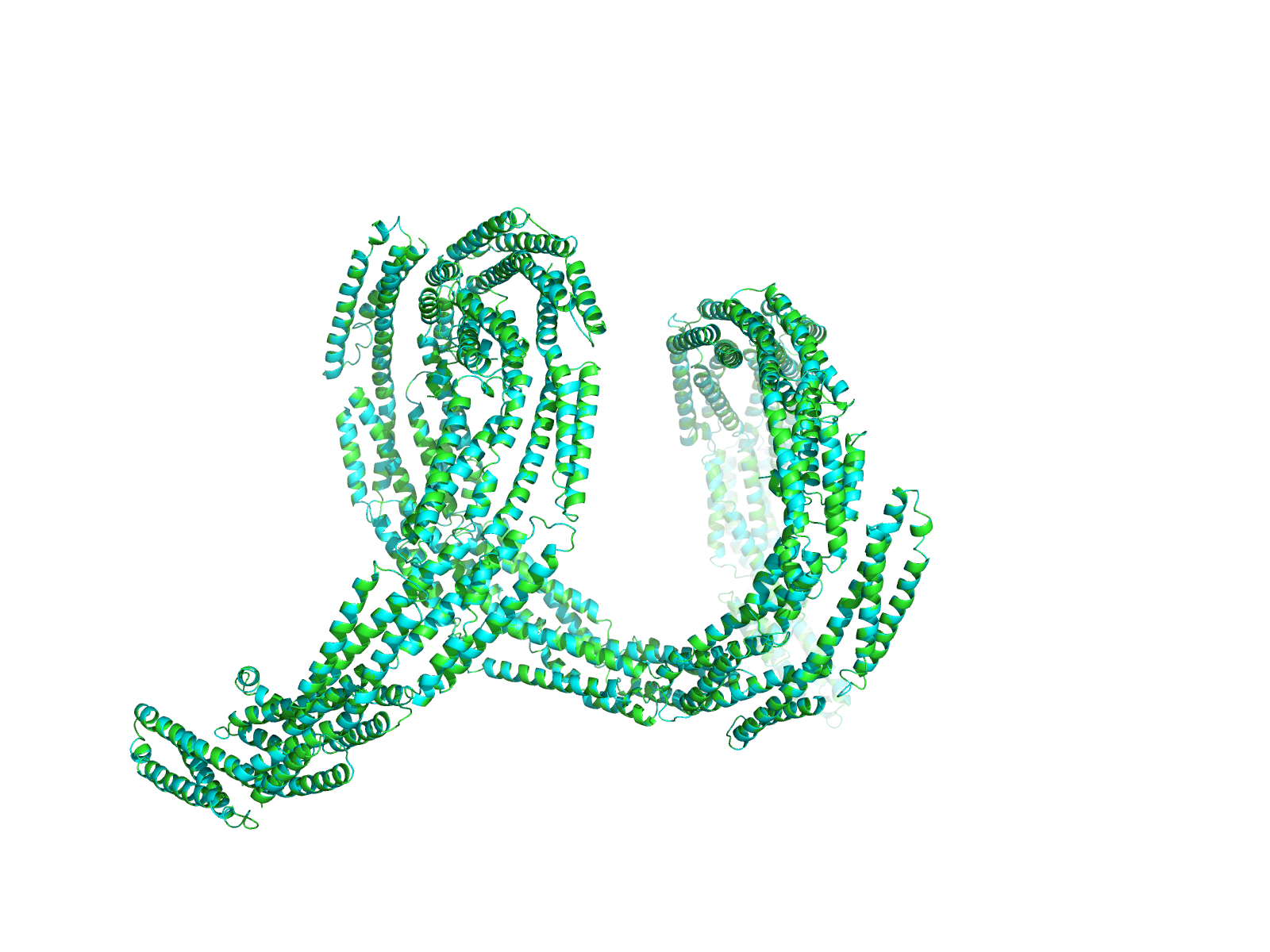
Bacteria spread through cell division; once a bacterial cell has grown to its full size, it starts to tighten a belt around its middle, and the belt grows ever tighter until it eventually severs the old cell into two separate cells. As this process is taking place, the cell has to make an exact copy of its genome, so that each new bacterial cell contains the same genetic information as all the others. In this way, bacteria ensure that they maintain their powerful defence mechanisms in each new cell as they spread through the body.

The work, published in Nature Communications, reveals that the atomic structure of the EzrA protein is as complex as it is beautiful. An unusual conformation, EzrA is shaped like a hoop, the middle of which is able to accommodate the belt-tightening protein, FtsZ. EzrA’s hoop-like shape prevents the FtsZ protein from escaping until it has fulfilled its purpose: causing the cell belt to contract and divide the cell in two whilst the genome of the parental cell is being copied and passed to the daughter.
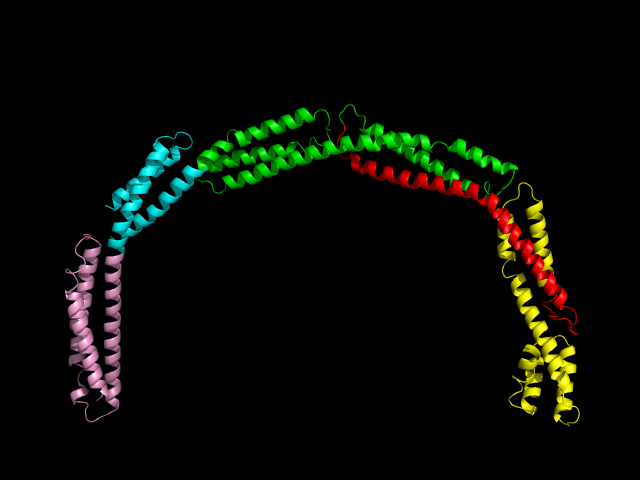
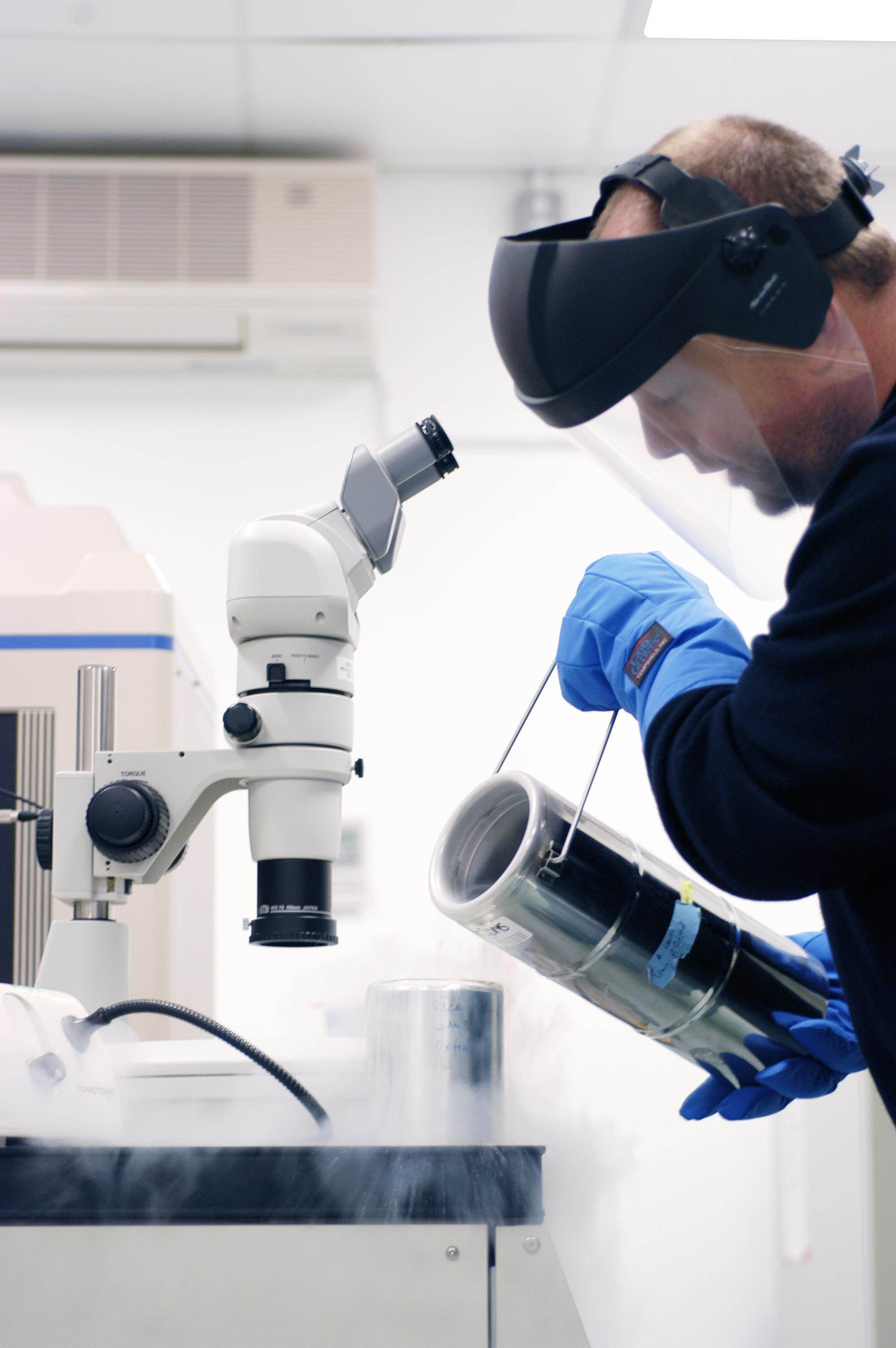
Research into the atomic structures of biological organisms is becoming ever more achievable as scientific technology advances. Powerful scientific machines like Diamond allow scientists to see things that are far too small to be glimpsed with a standard microscope. Over the course of its lifetime, Diamond has supported the solving of over 2000 protein structures such as this.
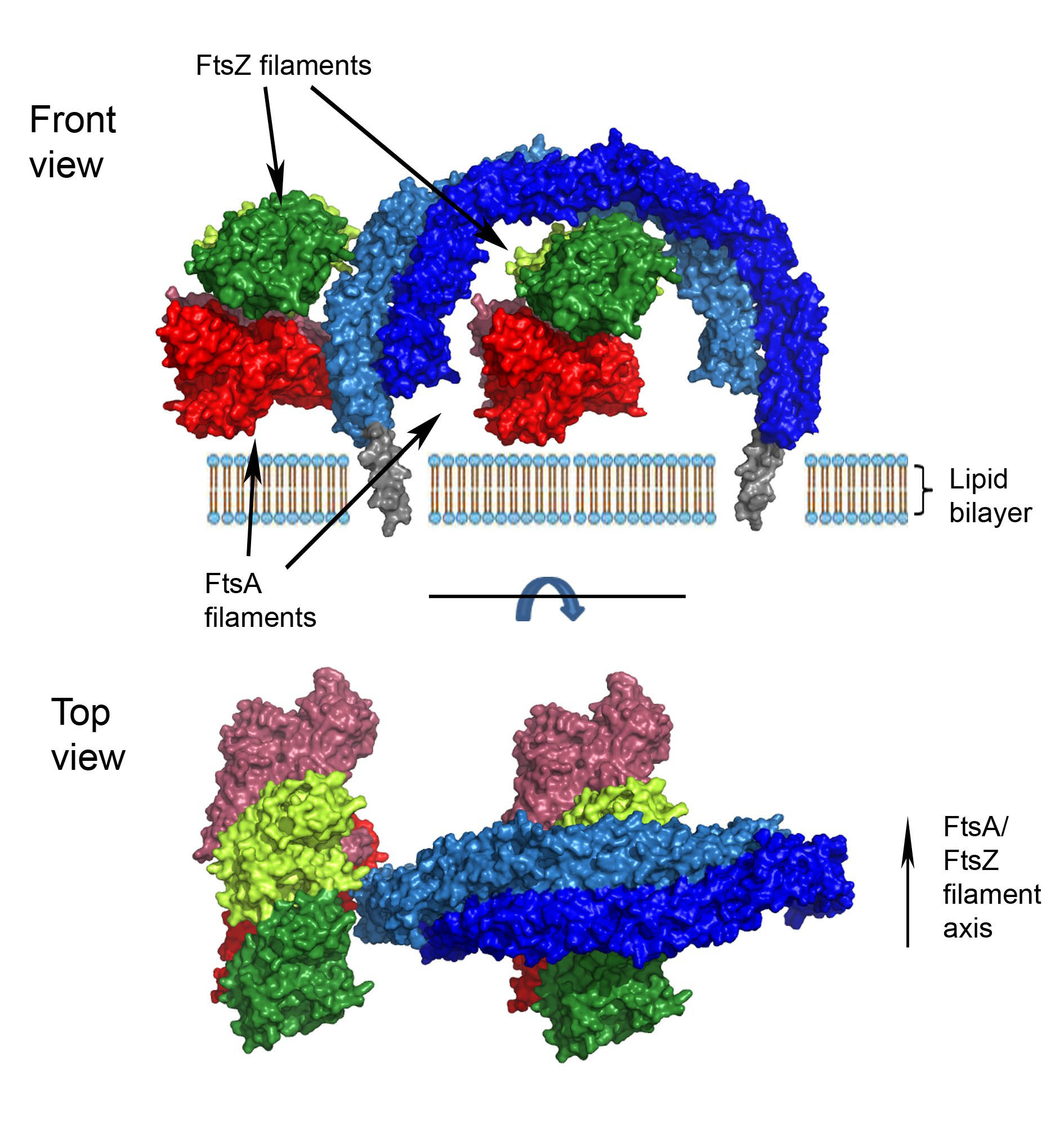
A model of how the EzrA protein, shown as a green molecular surface, interacts with the bacterial cytoskeletal proteins FtsZ (pink) and FtsA (blue). FtsZ is the belt that drives the constriction of the cell at the site of division and FtsA anchors the belt to the cell membrane.
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.